An oft-touted “fact” by creation skeptics is that young-earth creationists are “species fixists.” They claim creationists believe God originally created with present levels of biodiversity. On the other end of the spectrum are creation skeptics who label creationists as “hyper-evolutionists.” They imply that creationists are incapable of explaining the rapid diversification of animals that survived the Flood aboard Noah’s Ark. Both charges are, at best, naïve, and at worst, disingenuous.
The following is a summary of “The CRS eKINDS research initiative: Where we have been and where we are headed from here“ by Jean Lightner and Kevin Anderson, and of the surrounding discussion and research pertaining to it. The views expressed are not necessarily those of New Creation.
What is eKINDS?
Creationists actually believe that, in the beginning, God created distinct “kinds,” or “baramins,” of animals and plants. God created them with the ability to multiply, adapt, and form new species. The created kinds that survived the Flood diversified into the array of species we see today. Others we see preserved in the post-Flood fossil record. Baraminology, the study of created kinds, has flourished over the past three decades. Nevertheless, we still have unanswered questions that remain:
- Which organisms today descend from the same created kind?
- Which mechanisms are involved in creating the diversity that we see within created kinds?
- Can we trace the natural history of various animal kinds as they moved from the Ark and repopulated the earth?
In 2016, the Creation Research Society (CRS) founded the eKINDS research initiative. The goal of Examination of Kinds In Natural Diversification and Speciation is to resolve these very questions and more.
The Criteria of Baraminology
In a nutshell, baraminology uses suites of additive criteria that unite smaller groups of organisms, called monobaramins, into larger groups. Suites of subtractive criteria break up larger groups of organisms, called apobaramins, into smaller groups. Through successive approximation, the goal is to identify a group of organisms that is both a monobaramin and an apobaramin. This is called a holobaramin.
One of the most helpful criteria at a baraminologist’s disposal is hybridization. Reproduction involves such great complexities in every stage―from the union of sperm and egg to the birth of healthy offspring―that most baraminologists consider it only possible if the organisms in question were specifically designed to reproduce. Therefore, the ability to bring forth is a positive confirmation that two organisms belong to the same baramin. Baraminology studies that use the hybridization criterion show that baramin often approximate to the taxonomic rank of family, or even the order in some cases.
However, the hybridization criterion does have its handicaps. For any number of reasons, organisms can lose the ability to interbreed despite belonging to the same baramin. (e.g. To date, no one has interbred cheetahs with any other cat species.) Researchers also cannot readily apply it to non-captive animals or those that don’t naturally interbreed in the wild. To help with this, they have developed statistical baraminology tests like Baraminic Distance Correlation (BDC) and Multidimensional Scaling (MDS). These methods have also proven helpful, but they are not without limitations of their own.
A New Baraminological Tool
With the initiation of eKINDS came a new baraminology tool: whole-genome content analysis. This method utilizes molecular data that is accumulating for many species. Specifically, it measures the similarity in expressed orthologous protein content between species and assigns them to individual baramins. Researchers base it on two starting assumptions, the validity of which they can test with further research. One is that God created different baramins with a unique array of protein coding genes that remained relatively unchanged throughout history. The second assumption is that each baramin has, by and large, retained what makes them distinct from other baramin.
The First eKNIDS Experiment
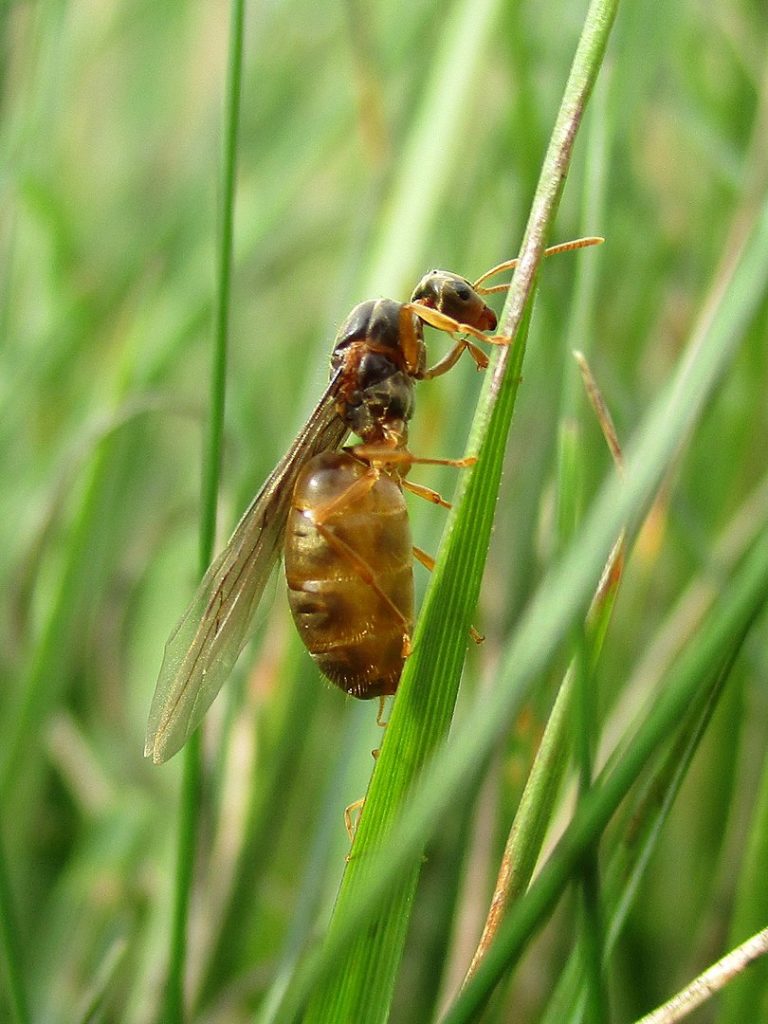

By Dat doris – Own work, CC BY-SA 4.0, https://commons.wikimedia.org/w/index.php?curid=92465587
The first group of animals on which the eKINDS project used this method consisted of 104 species of insects. They were from the orders Diptera (flies), Hemiptera (“true bugs”), Hymenoptera (wasps, bees, ants, etc), and Lepidoptera (moths and butterflies). While the results are as of yet preliminary, they do display some interesting patterns:
- Diptera formed two tentative groups. The first consists of black flies, gnats, crane flies, midges, and mosquitoes. The second included horse flies and fruit flies.
- Hemiptera divided into two tentative groups, but only one of them was statistically significant.
- The ants, bees, wasps, and other species of Hymenoptera included in this analysis fell into one group.
- The ten species of Lepidoptera included in this study represent six different superfamilies. Despite this, they all formed one group.
Comparing Humans and Primates
The researchers also used whole-genome content analysis to compare humans and other primates with other mammals and birds. The three species of humans for which we have genomic data were Homo sapiens (us), Neanderthals, and Denisovans. Researchers found that these formed a group clearly divided from chimpanzees and other great apes, which were more aligned with Old World and New World monkeys. Even more so, they found that the differences between humans and other primates were comparable to the differences between humans and some cats!
Heat Map from Lightner et al, 20181
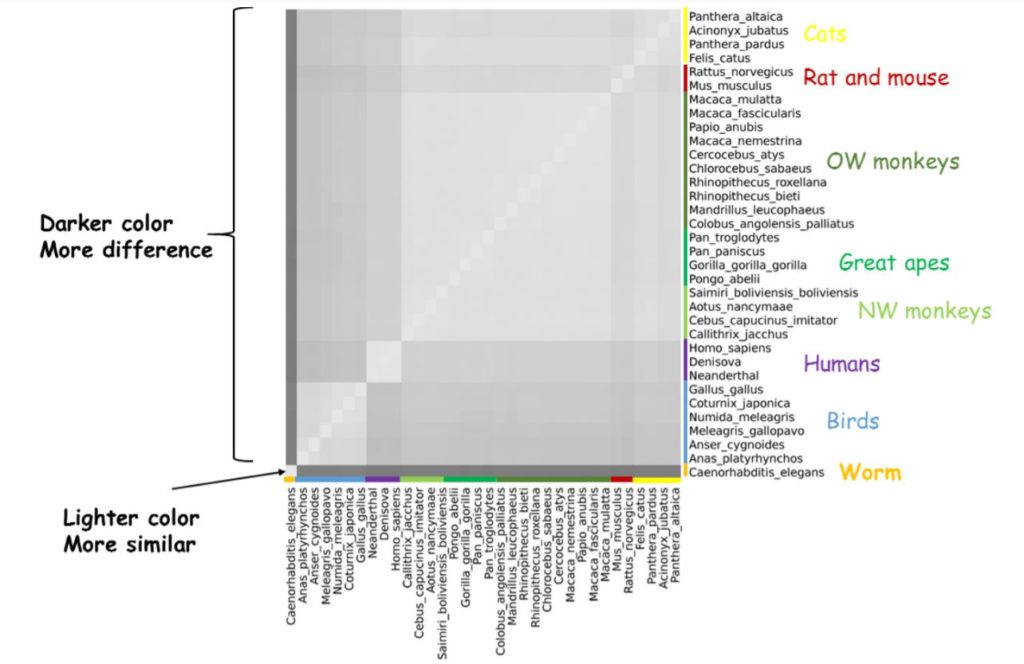
From Lightner et al, 2018.2
What Do the Results Tell Us?
The clear discontinuities among insects and between humans and all other mammals are not a predicted result of universal common ancestry. That would maintain that all of biodiversity forms a gradient. It is, however, very consistent with the creationist predictions that discontinuities should punctuate the Tree of Life, reflecting different baramins.
They suggest that it “is fully consistent with the belief that humans were separately created in the image of God, and should not be too surprising given that a significant number of genes are either unique to humans or distinct in humans.”
If baraminology has taught us anything, it is that there can be great diversity within a baramin. But how did all of this biodiversity come about? One common creationist explanation is that post-Creation changes within a baramin occurred as a result of natural selection, mutation, and the recombination of created alleles. There is no reason to doubt that these were factors in operation in the past as now. At the same time, however, there is evidence that other biodiversification mechanisms were also at work.
Future Scientific Inquiry
One mechanism that participants of the eKINDS project are eager to look into further is DNA editing. Activation-induced cytidine deaminase (AID) is a known enzyme used in immune system DNA editing to create diverse antibodies. This process involves somatic hypermutation and class switch recombination. Could AID or a similar enzyme also have played a role in generating adaptive mutations in vertebrates that resulted in heritable genetic changes? This has not yet been demonstrated, but provides an excellent launch pad for future scientific inquiry.

By Viksah626 – Own work, CC BY-SA 4.0, https://commons.wikimedia.org/w/index.php?curid=49477133
Another factor that eKINDS investigates is the importance of the founder effect. This refers to a decrease of available genetic variation when a new population of organisms form and establish from a small number of individuals from a larger population. As animals multiplied and spread across the earth’s surface after the Flood, they would have been spoiled for choice over the new, empty environments they could colonize. This factor alone would produce a number of founder effects. It could be a key ingredient in a recipe for rapid biodiversification, largely untapped by creationists as of yet.
Conclusion
There is still much to learn about baraminic membership and the mechanisms involved in their diversification in the pre-Flood and post-Flood world. Currently, the eKINDS researchers are carrying out an in-depth investigation of the kingfishers (Alcedinidae) and landfowl (Galliformes) to see what they learned of their natural history.
The authors conclude that “the eKINDS project has already helped to expand the creation model regarding diversification and adaptation of the original kinds. It is our hope that, by the grace of God, the project will gain momentum and more researchers will contribute. This would enable us, as believers, to significantly advance the creation model as we come to more fully understand the world God created.”
Footnotes
- Lightner, J.K., and K. Anderson. (2018). “The CRS eKINDS research initiative: Where we have been and where we are headed from here.” In Proceedings of the Eighth International Conference on Creationism, ed. J.H. Whitmore, pp. 185–190. Pittsburgh, Pennsylvania: Creation Science Fellowship. ↩︎
- Lightner, 2018, pp. 185–190. (Footnote 1) ↩︎